To get started and become familiar with the types of analyses you can perform using SpatialRNA, we recommend going through the first three tutorials before scaling up your analysis to larger sets of samples.
Please refer to the code repository for all scripts and defined snakemake rules for reproducing the results here. In this notebook, we plot the transcript-based niches.
Processing steps
Preparing input transcripts
Filter out unwanted transcripts (e.g., control probes or low quality, such as
qv < 20in Xenium data).Save the cleaned list of transcripts (with spatial coordinates columns) to:
./data/MySampleNameX/raw/MySampleNameX.csvFor multiple samples, save them using the same structure.
Generate tiled spatial RNA graphs
See 01
_run _build _tile _graph _and _train, which defines two rules: generate_graph— Runs./code/run_build_graph.py, which uses theSpatialRNAfunction to tile the input tissue into 100 tiles. Each Snakemake job processes one tile.run_on_disk_train_on_all_tilesing— Runs./code/run_train_with_myod.py, which builds anOnDiskDataLoaderto load tile graphs in shuffled batches, then trains a selected GNN model.
Embedding inference with the trained model
See 02
_run _infer _and _cluster, which defines: run_infer— Runs inference withSpatialRNAto generate embeddings for transcripts in individual tiles.run_gmm_cluster— Clusters all transcripts jointly using Gaussian Mixture Model.
Embedding inference for tiles can be parallelised across Snakemake jobs.
Visualisation of the transcript-based molecular niches¶
import pandas as pd
import numpy as np
import os.path as osp
import time
import random
import sys
import matplotlib.pyplot as plt
from pathlib import Path
ovc_df = pd.read_csv("../../../case_study_xenium_5k_panel/data/OvarianCancer/raw/OvarianCancer.csv",
usecols=["x_location","y_location"])
out_dir = "../../../case_study_xenium_5k_panel_revision/output/on_disk_train_on_all_tiles/GraphSAGE_trained/nnedge10000_seed20251/on_disk_batch_size3/gmm_clusters/gmm15_trained//"
all_gmm_res_k15 = [pd.read_csv(f"{out_dir}/OvarianCancer_tile{x}.txt",names=["gmm15"]) for x in range(0,100)]
out_dir = "../../../case_study_xenium_5k_panel_revision/output/on_disk_train_on_all_tiles/GraphSAGE_trained/nnedge10000_seed20251/on_disk_batch_size3/gmm_clusters/gmm10_trained/"
all_gmm_res_k10 = [pd.read_csv(f"{out_dir}/OvarianCancer_tile{x}.txt",names=["gmm10"]) for x in range(0,100)]
all_gmm_res_k15 = pd.concat(all_gmm_res_k15)
all_gmm_res_k10 = pd.concat(all_gmm_res_k10)
all_gmm_res_id = [pd.read_csv(f"../../../case_study_xenium_5k_panel_revision/data/OvarianCancer/embedding/OvarianCancer_data_tile{x}input_tx_id.csv") for x in range(0,100)]
all_gmm_res_id = pd.concat(all_gmm_res_id)
all_gmm_res_id.shape,all_gmm_res_k10.shape,all_gmm_res_k15.shape,ovc_df.shape
((106423555, 1), (106423555, 1), (106423555, 1), (106423555, 2))ovc_df.head(5)
ovc_df = ovc_df.loc[all_gmm_res_id["tx_id"]]
ovc_df["gmm10"] = all_gmm_res_k10["gmm10"].astype("str").values
ovc_df["gmm15"] = all_gmm_res_k15["gmm15"].astype("str").values
ovc_df.head(5)
from spatialrna import viz
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from matplotlib.colors import ListedColormap
colors = ['#e6194b', '#3cb44b', '#ffe119', '#4363d8', '#f58231',
'#911eb4', '#46f0f0', '#f032e6', '#bcf60c', '#fabebe', '#008080','#e6beff',
'#9a6324', '#fffac8', '#800000', '#aaffc3', '#808000', '#ffd8b1',
'#000075', '#808080', '#ffffff', '#000000']
len(colors)
22customise_cmap_k10 = dict(zip([str(x) for x in range(10)],colors))
customise_cmap_k15 = dict(zip([str(x) for x in range(15)],colors))
import matplotlib.colors as mcolors
ovc_df.shape
(106423555, 4)GMM 15, overall across tissue¶
GMM 15 hex bin plot¶
import matplotlib.pyplot as plt
plt.rcParams['figure.facecolor'] = 'black'
plt.style.use('default')
p_fig,ax = viz.plot_hex_bin(tx_meta = ovc_df.sample(50000000),
x="x_location",
y="y_location",
cluster_labels="gmm15",
bin_width=5,
bin_thresh=5,
cmap=customise_cmap_k15,
background_color="white",dpi=300,figsize=(10,10))
#ax.invert_yaxis()
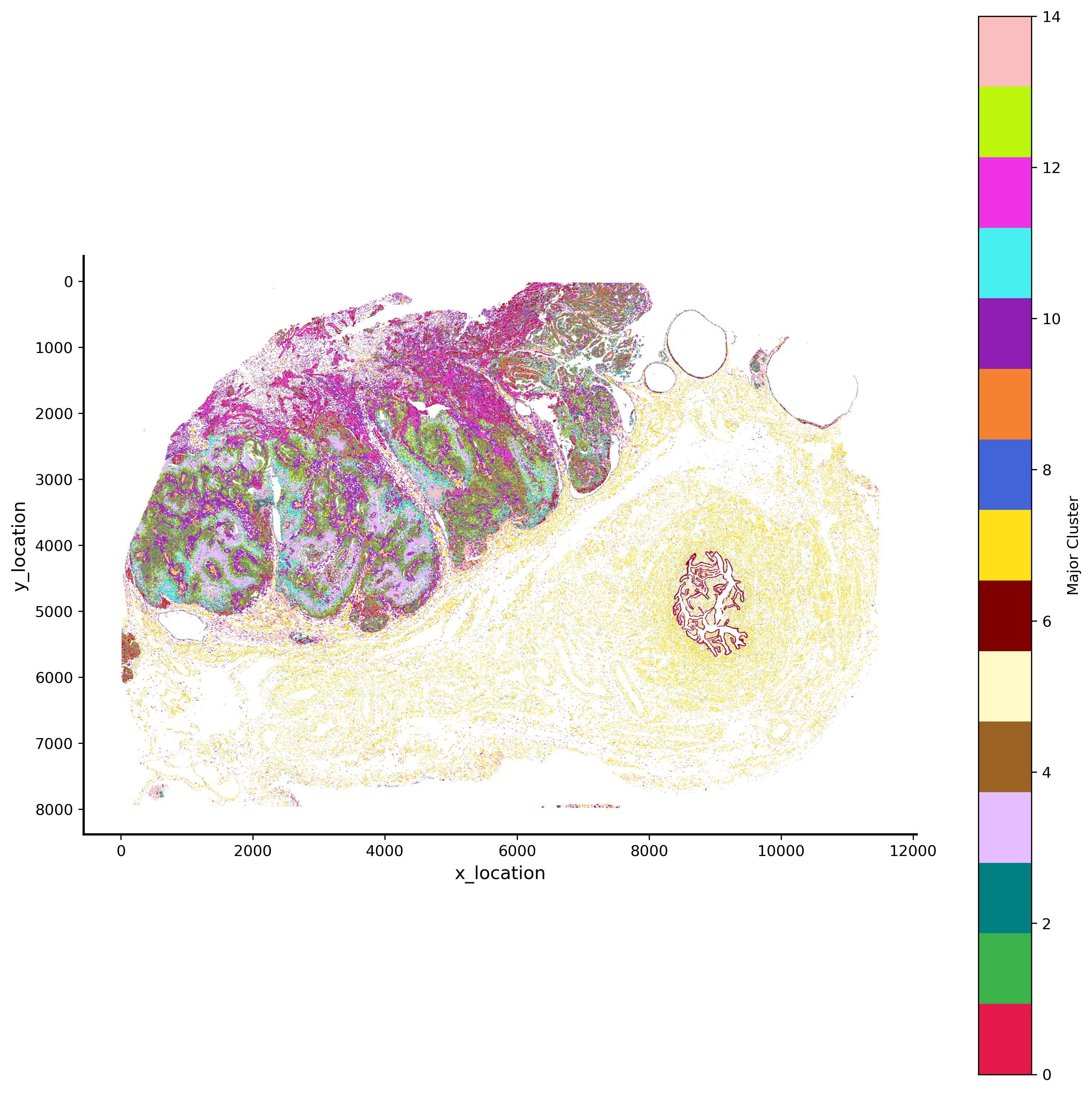
GMM 15 pixel plot¶
import matplotlib.pyplot as plt
plt.rcParams['figure.facecolor'] = 'black'
plt.style.use('dark_background')
# p_fig,ax = viz.plot_pixel(tx_meta = ovc_df.sample(50000000),
# x="x_location",
# cluster_labels="gmm15",
# y="y_location",
# pixel_size=0.5,min_points=1,
# join_method="avg",
# cmap=customise_cmap_k15,
# background_color="black",dpi=300,figsize=(10,10))
#ax.invert_yaxis()
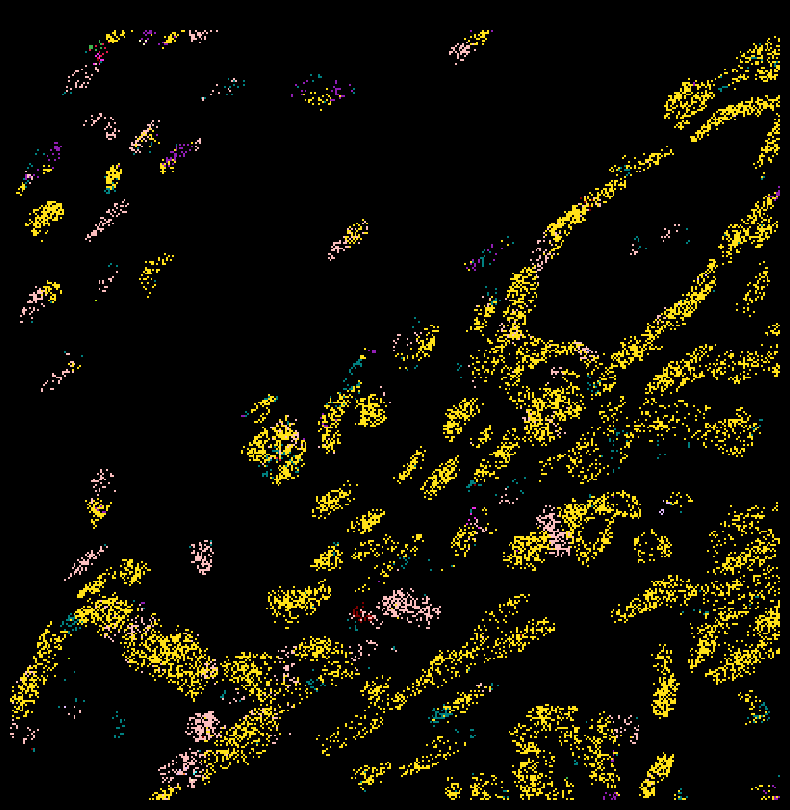
Zoom in¶
subset_df = ovc_df[(ovc_df["x_location"] < 500) & (ovc_df["y_location"] < 4000) ]
plt.rcParams['figure.facecolor'] = 'black'
fig,ax = viz.plot_pixel(tx_meta =
ovc_df[(ovc_df["x_location"] > 4800) & (ovc_df["x_location"] < 5000) &
(ovc_df["y_location"] < 5000) & (ovc_df["y_location"] > 4800) ],
x="x_location",
cluster_labels="gmm15",
y="y_location",
pixel_size=0.5,
min_points=1,
cmap=customise_cmap_k15,
join_method="major",
background_color="black",dpi=100,figsize=(10,10))
ax.invert_yaxis()
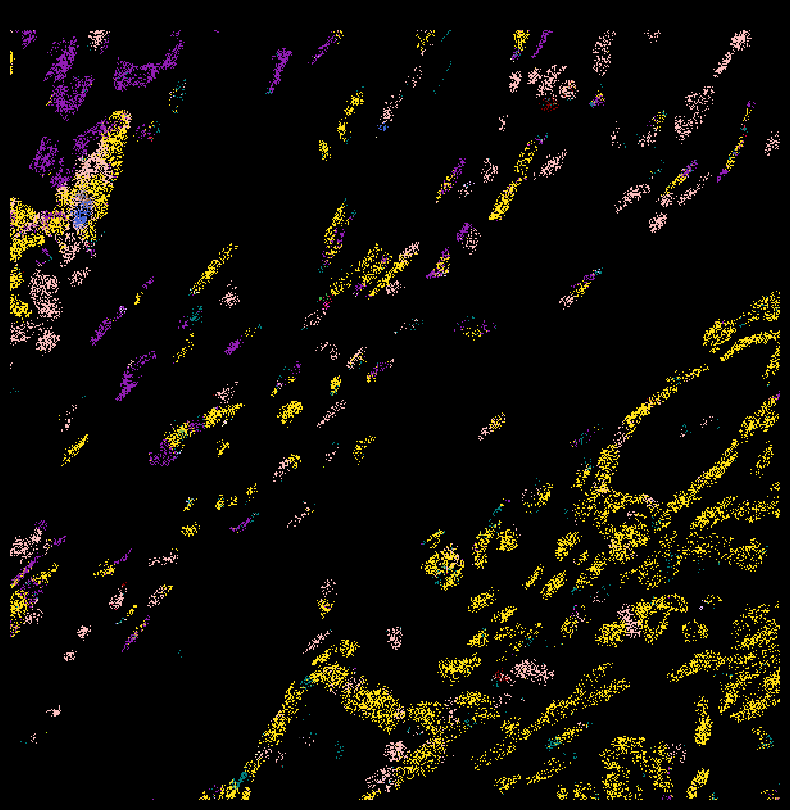
plt.rcParams['figure.facecolor'] = 'black'
fig,ax = viz.plot_pixel(tx_meta =
ovc_df[(ovc_df["x_location"] > 4700) & (ovc_df["x_location"] < 5000) &
(ovc_df["y_location"] < 5000) & (ovc_df["y_location"] > 4700) ],
x="x_location",
cluster_labels="gmm15",
y="y_location",
pixel_size=0.5,
min_points=1,
cmap=customise_cmap_k15,
join_method="avg",
background_color="black",dpi=100,figsize=(10,10))
ax.invert_yaxis()
plt.rcParams['figure.facecolor'] = 'black'
fig,ax = viz.plot_pixel(
tx_meta = ovc_df[(ovc_df["x_location"] > 500) & (ovc_df["x_location"] < 1000) &
(ovc_df["y_location"] < 4000) & (ovc_df["y_location"] > 3500) ],
x="x_location",
cluster_labels="gmm15",
y="y_location",
pixel_size=0.5,
min_points=1,
join_method="avg",
cmap=customise_cmap_k15,
background_color="black",dpi=100,figsize=(10,10))
ax.invert_yaxis()
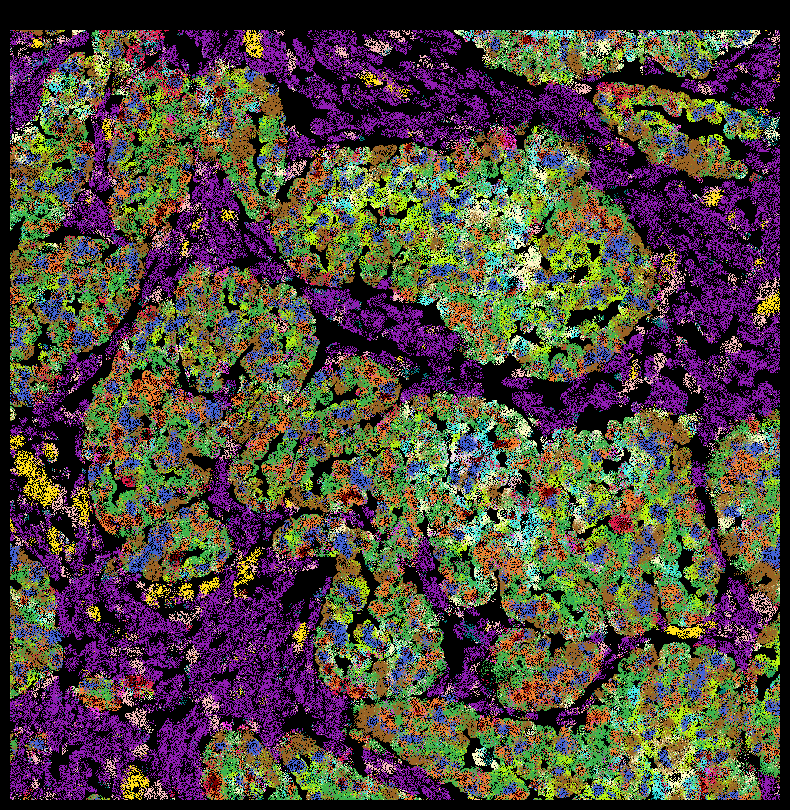
Save to file instead of plotting¶
viz.plot_pixel(
tx_meta = ovc_df[(ovc_df["x_location"] > 500) & (ovc_df["x_location"] < 1000) &
(ovc_df["y_location"] < 4000) & (ovc_df["y_location"] > 3500) ],
x="x_location",
cluster_labels="gmm15",
y="y_location",
pixel_size=0.5,
min_points=1,
cmap=customise_cmap_k15,
join_method="avg",
output_path = "../figures/crop1_5k.png",
background_color="black"
)
Saved pixelated RGB image to ../figures/crop1_5k.png
15 molecular niches for whole tissue¶
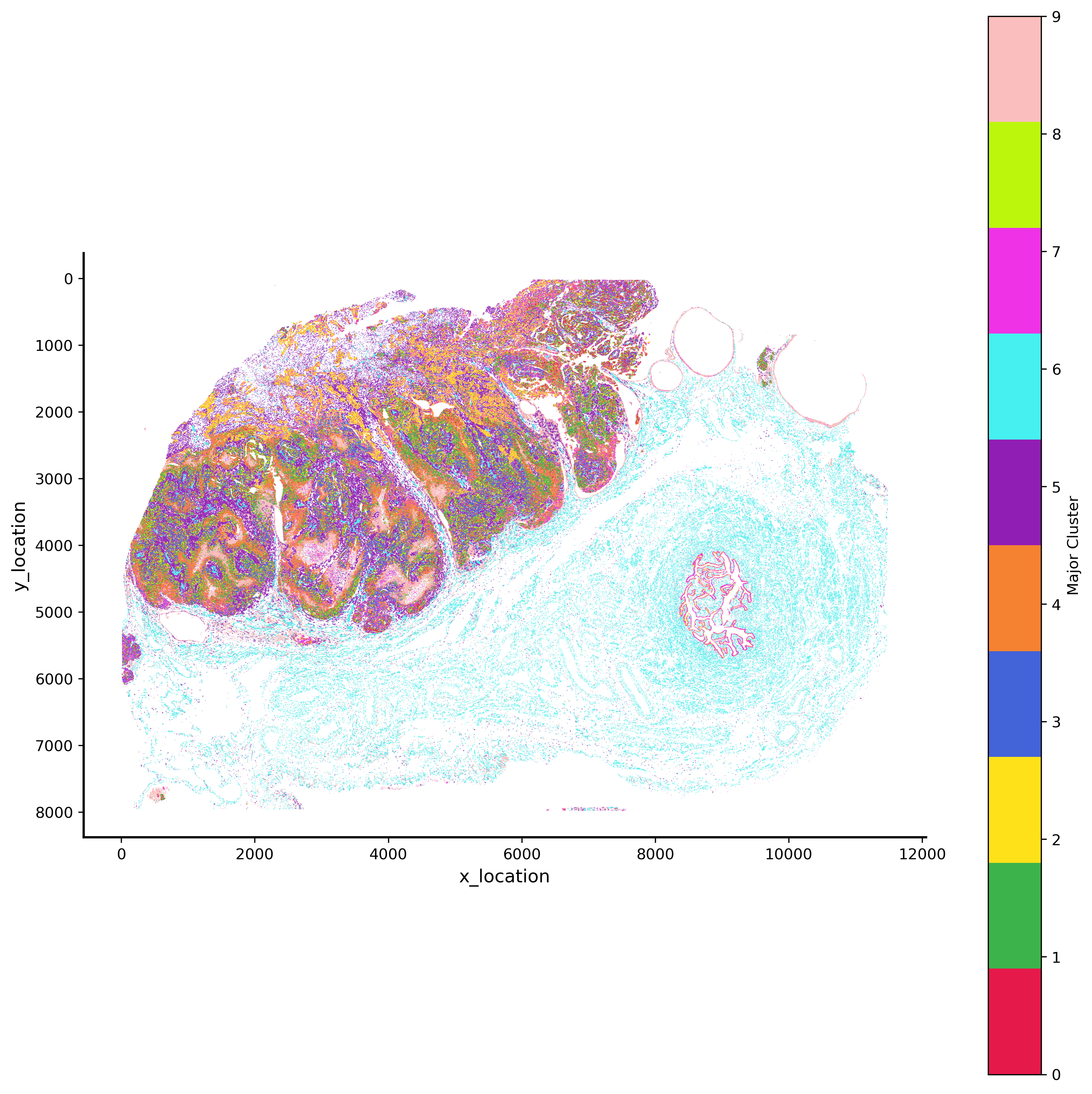
GMM 10 hex bin plot¶
import matplotlib.pyplot as plt
plt.rcParams['figure.facecolor'] = 'black'
plt.style.use('default')
p_fig,ax = viz.plot_hex_bin(tx_meta = ovc_df.sample(50000000),
x="x_location",
y="y_location",
cluster_labels="gmm10",
bin_width=5,
bin_thresh=5,
cmap=customise_cmap_k10,
background_color="white",dpi=300,figsize=(10,10))
#ax.invert_yaxis()
The history saving thread hit an unexpected error (OperationalError('attempt to write a readonly database')).History will not be written to the database.